library(here)
library(terra)
library(sf)
library(lubridate)
library(picMaps)
library(rnaturalearth)
library(reticulate)
library(tidyverse)
library(foreach)
library(foreach)
library(future)
library(doFuture)
library(progressr)
library(ggplot2)
library(ggspatial)
localwd <- here("_R_code","task_5_download_prediction_data")
copernicus_data <- file.path(localwd, "copernicus_data")
pred_dir <- file.path(localwd, "output")
if(!dir.exists(copernicus_data)) dir.create(copernicus_data)
if(!dir.exists(pred_dir)) dir.create(pred_dir)
reticulate::use_virtualenv("cm_py", required = TRUE)
cm <- reticulate::import("copernicusmarine")
lg <- reticulate::import("logging")
lg$getLogger("copernicusmarine")$setLevel('WARN')Download Environmental Variables for Density Prediction
Note!!! Currently this is the way that our team downloaded oceanographic model data from Copernicus Marine Service. This more complicated method was necessary because the CopernicusMarinepackage did not have the ability to download ocean product data subsets. However, as of September 2025 this capability was repaired. So, the CopernicusMarine package is probably a more efficient way to download the data, and we will investigate this further.
The procedure for downloading the oceanographic prediction data is nearly identical to that in step 3. The only difference is specifying the spatial and temporal grid defined by the central Pacific study area and the tri-daily grid of days from July 1 through December 31 in the years 2017 and 2020-2024.
Packages and Python preparation
Defining the central Pacific
cenpac <- picMaps::cenpac()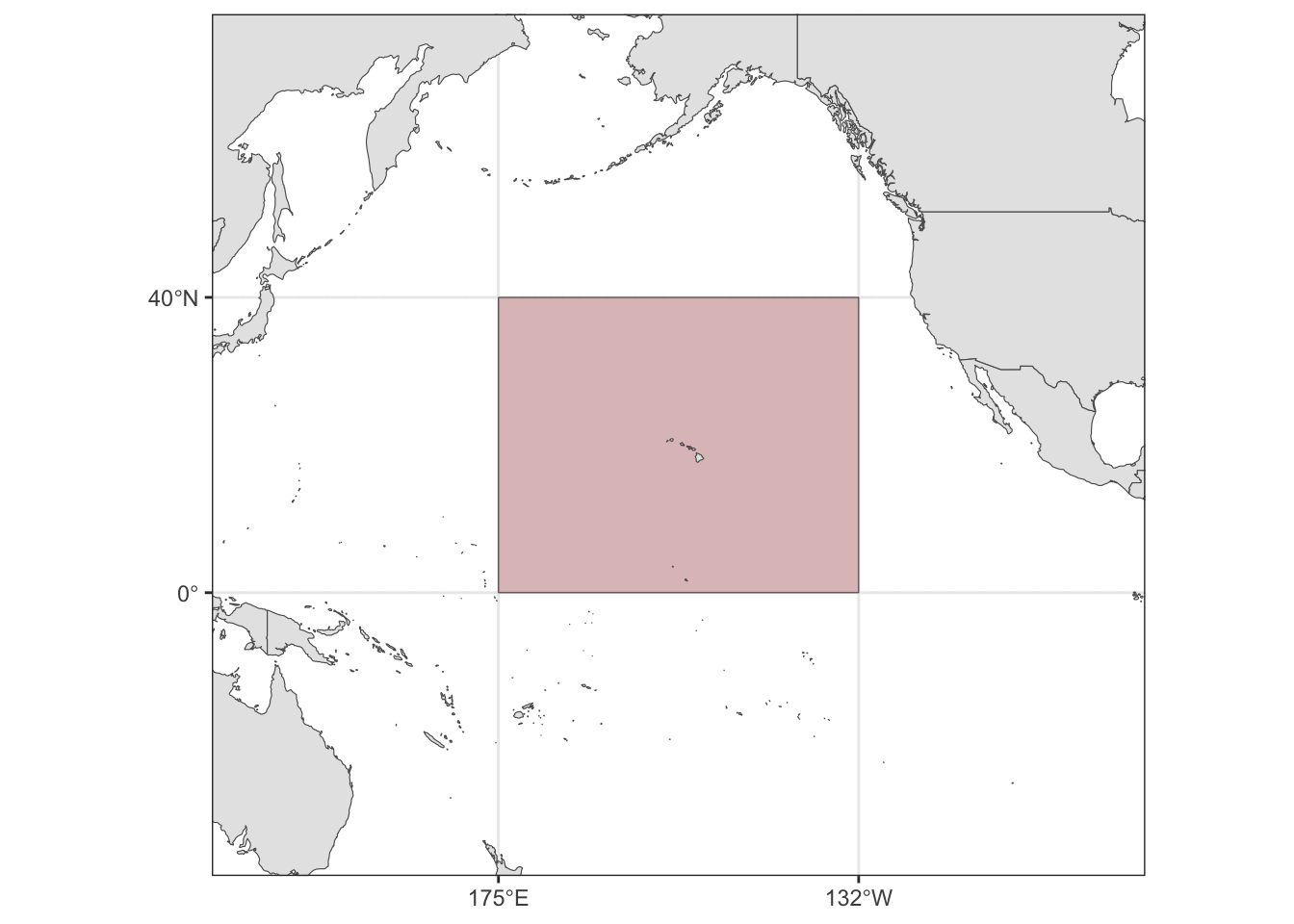
Downloading the environmental variables
years <- c(2017, 2020:2024)
bound <- sf::st_bbox(cenpac)
with_progress({
pb <- progressor(along = years)
pred_download <- foreach(i = seq_along(years),
.options.future = list(seed = TRUE)) %do%
{
time_span <- paste(years[i],c("-07-01","-12-31"), sep="")
nc_dl <- cm$subset(
dataset_id = "cmems_mod_glo_phy_my_0.083deg_P1D-m",
variables = as.list(c("mlotst","so","thetao","zos")),
minimum_longitude = bound$xmin - 1/6,
maximum_longitude = bound$xmax + 1/6,
minimum_latitude = bound$ymin - 1/6,
maximum_latitude = bound$ymax + 1/6,
start_datetime = time_span[1],
end_datetime = time_span[2],
minimum_depth = 0.5,
maximum_depth = 0.5,
coordinates_selection_method = "outside",
dataset_version = "202311",
disable_progress_bar = FALSE,
skip_existing = TRUE,
output_directory = copernicus_data
)
pb()
data.frame(year=years[i], filename=nc_dl$filename)
}
})
# plan("sequential")
pred_download <- do.call(rbind, pred_download)
write.csv(pred_download, file.path(copernicus_data, "pred_download.csv"), row.names = FALSE)Processing the raw netCDF files
# localwd <- here("_R_code","task_5_download_prediction_data")
# copernicus_data <- file.path(localwd, "copernicus_data")
# pred_dir <- file.path(localwd, "output")
# pred_download <- read.csv(file.path(copernicus_data, "pred_download.csv"))
years <- pred_download$year
with_progress({
pb <- progressor(along = years)
for(i in seq_along(years)){
filenm <- pred_download$filename[i]
env <- terra::rast(file.path(copernicus_data, filenm))
vnms <- terra::varnames(env)
new_vnms <- c("mld","salinity","sst","ssh")
dates <- time(env) |> unique() |> sort()
subs_idx <- which(rep(1:3, length=length(dates))==1)
out_env <- NULL
for(j in seq_along(vnms)){
tmp_env <- toMemory(env[vnms[j]])
names(tmp_env) <- rep(new_vnms[j],nlyr(tmp_env))
tmp_env <- tmp_env[[subs_idx]]
tmp_env <- terra::project(tmp_env, "+proj=longlat +ellps=WGS84 +lon_wrap=180 +datum=WGS84 +no_defs")
tmp_env <- terra::focal(tmp_env, 3, mean, na.policy="only", na.rm=TRUE) # Make sure there are no cells with missing data
out_env <- c(out_env, tmp_env)
}
out_env <- do.call(c, out_env)
# add focal sd
for(j in seq_along(vnms)){
tmp_env <- out_env[[names(out_env)==new_vnms[j]]]
tmp_env_sd <- focal(tmp_env, 3, sd, na.rm=TRUE)
names(tmp_env_sd) <- paste(names(tmp_env_sd),"sd",sep="_")
out_env <- c(out_env,tmp_env_sd)
}
writeRaster(out_env, file.path(pred_dir, paste0("cenpac_",years[i], "_env.tif")), overwrite=TRUE)
pb()
}
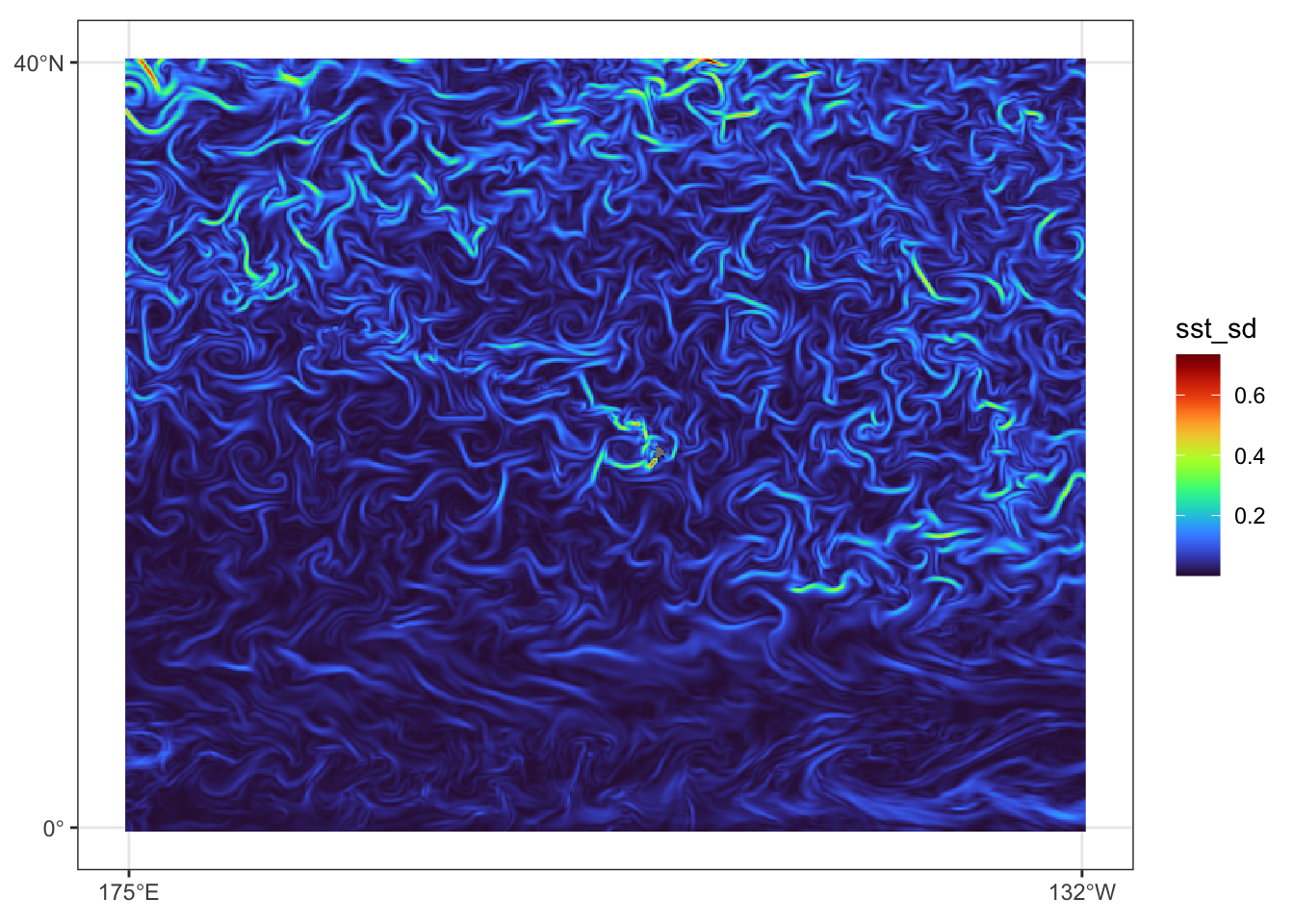
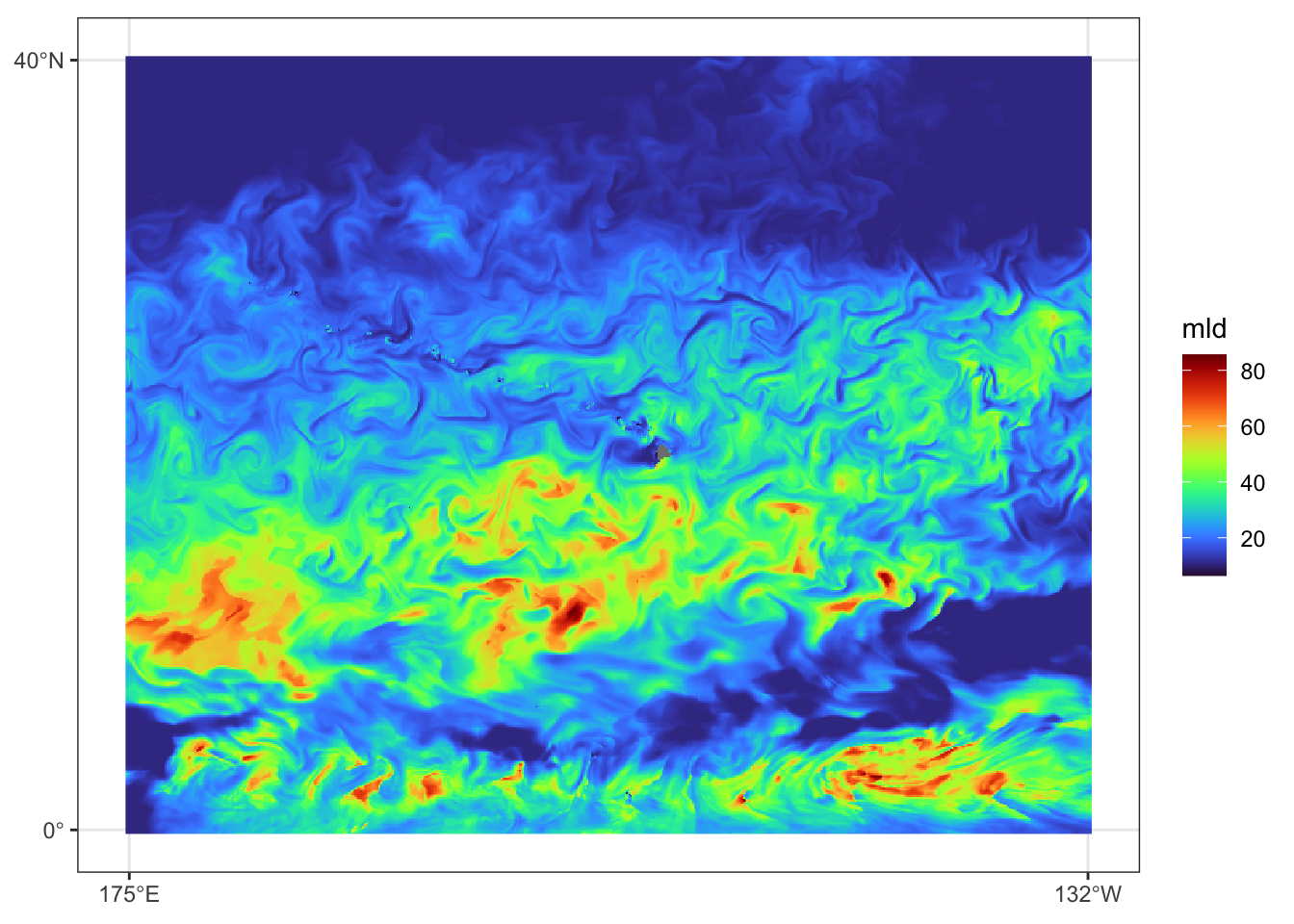
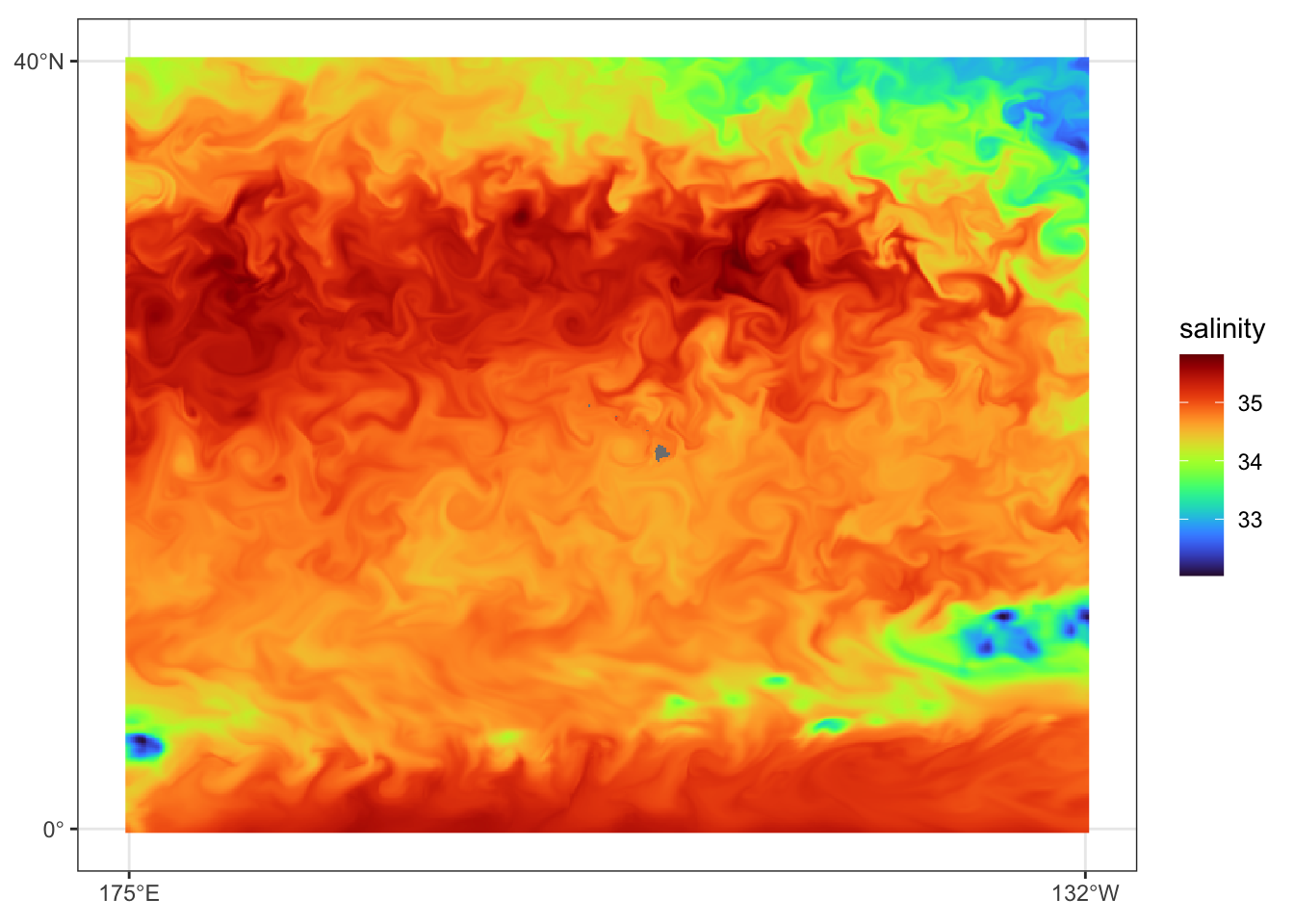
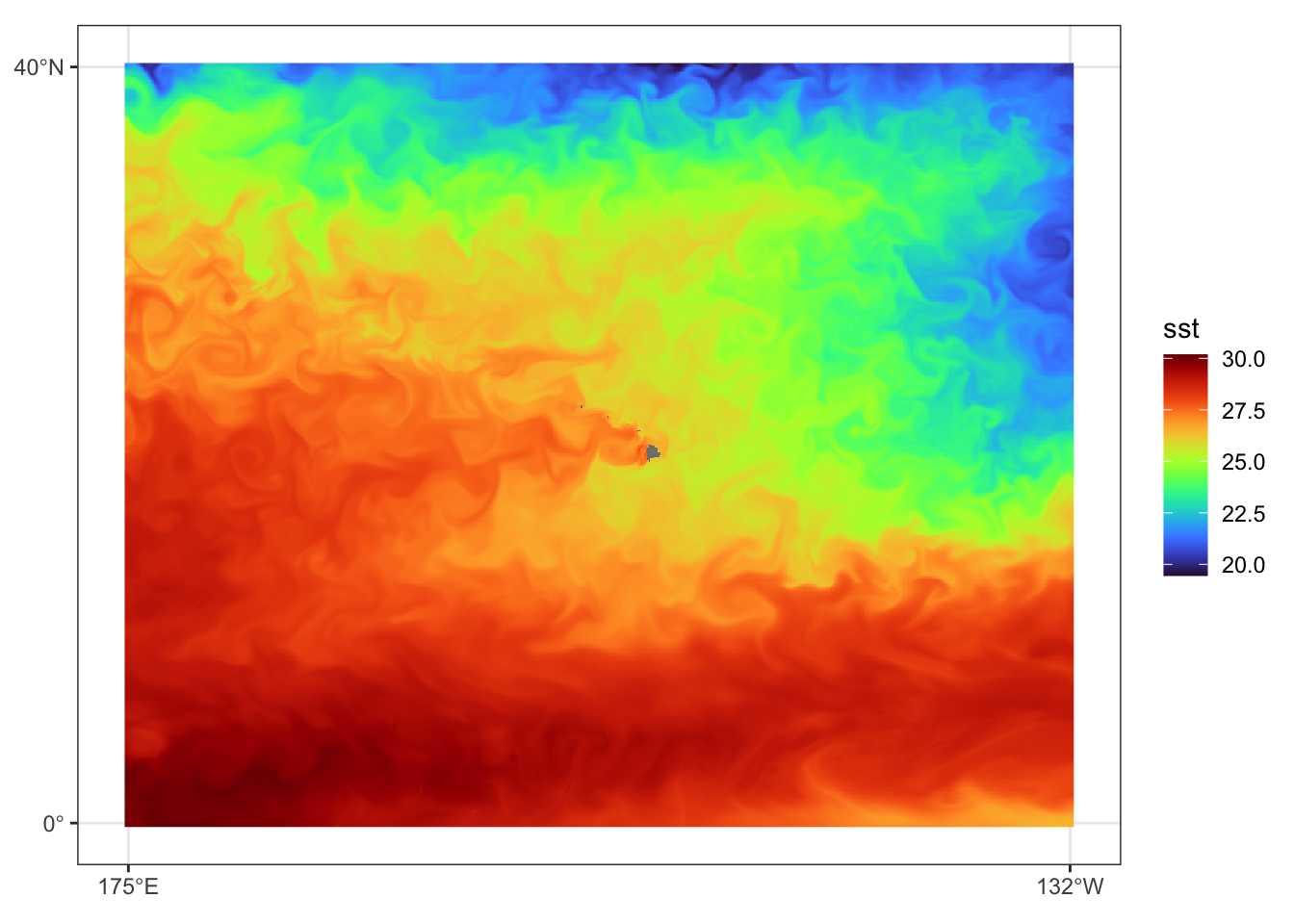
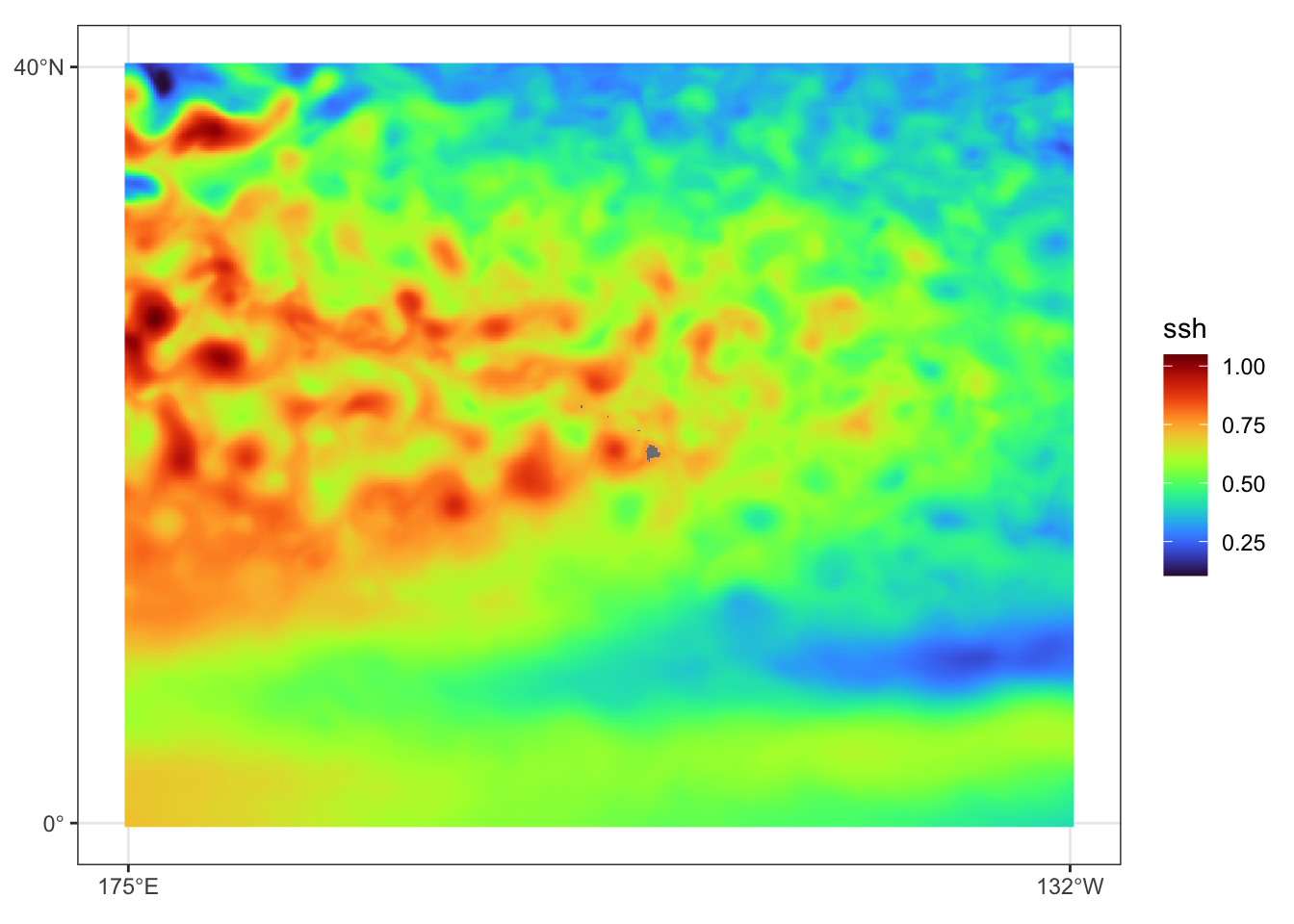
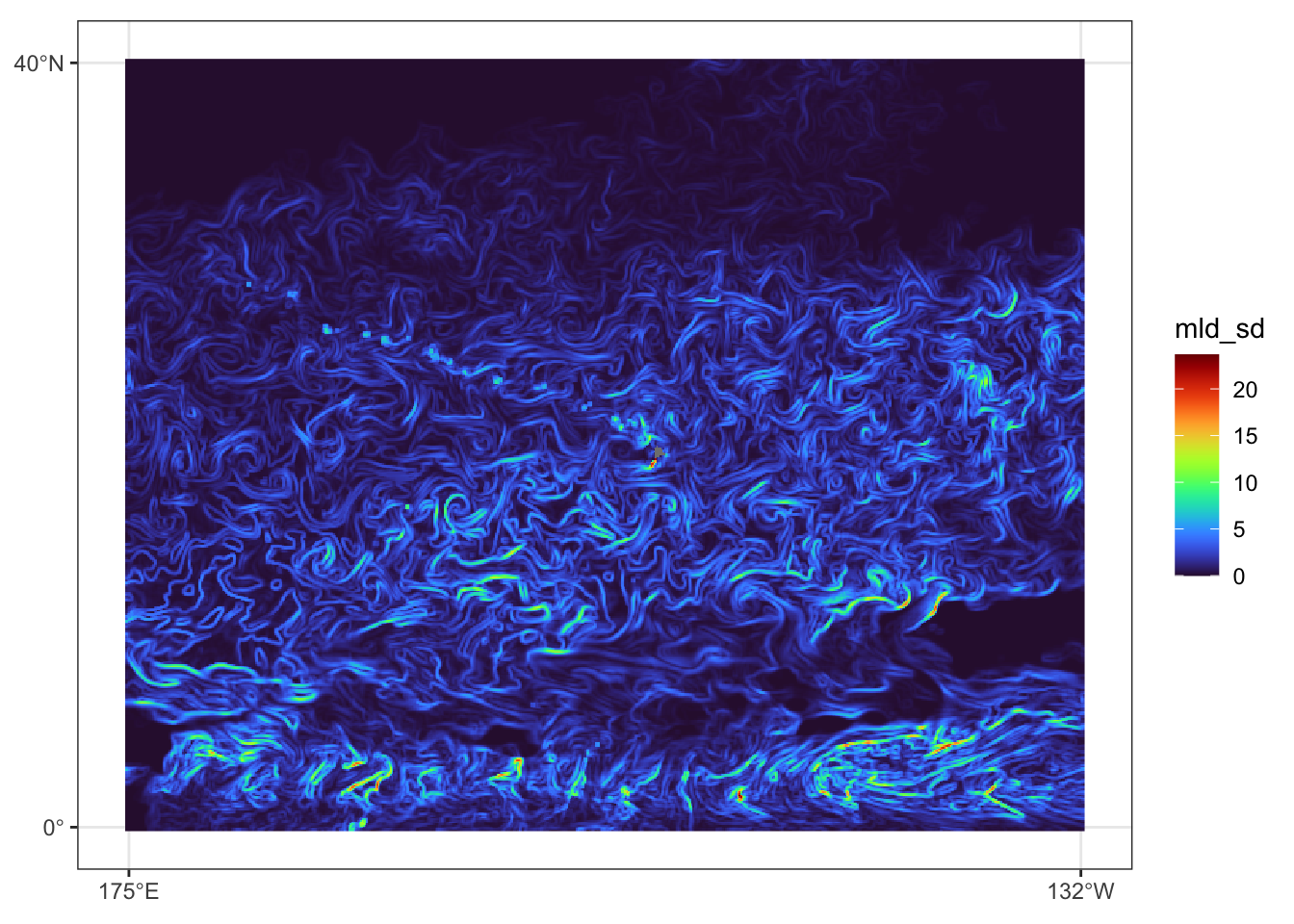
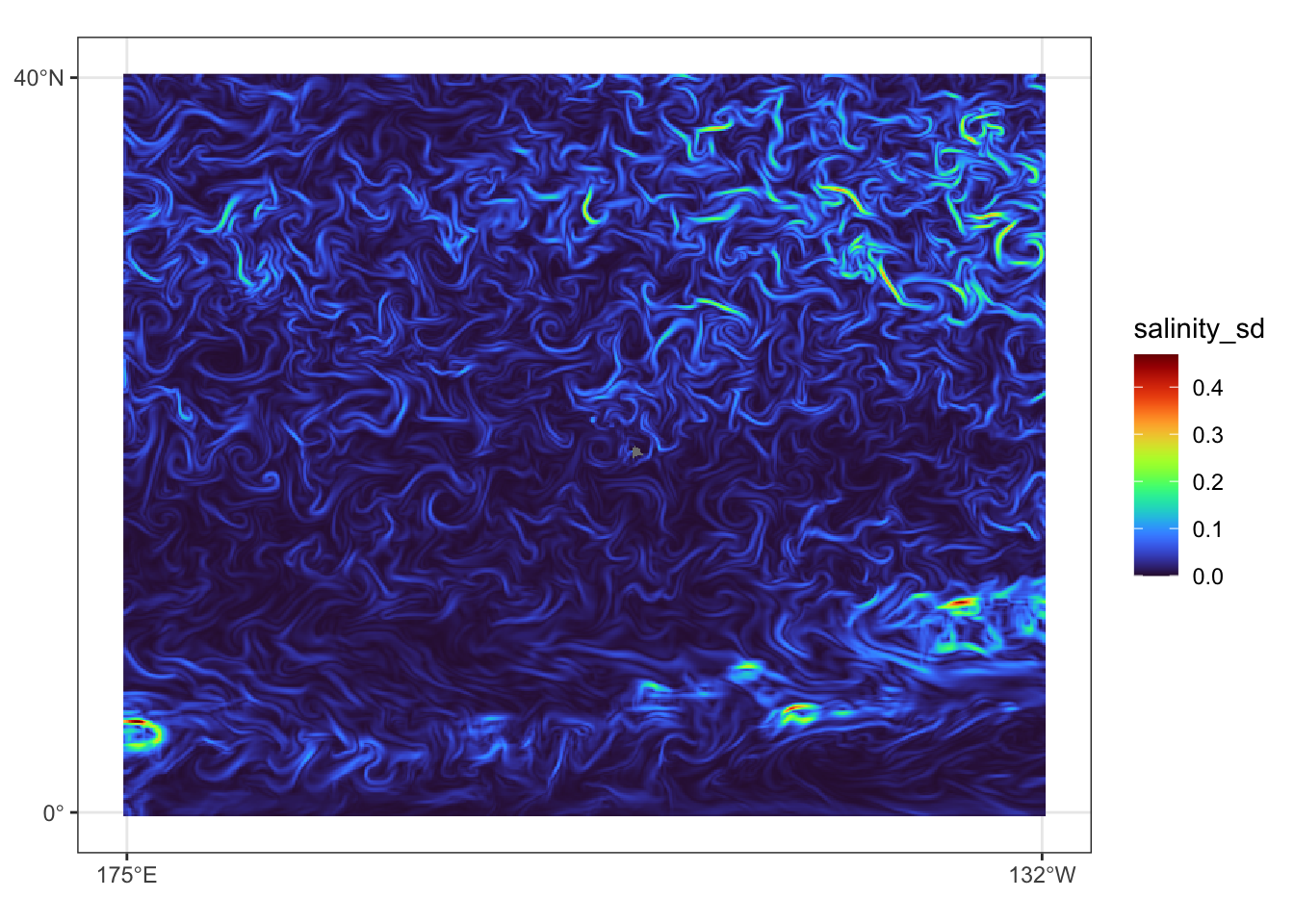
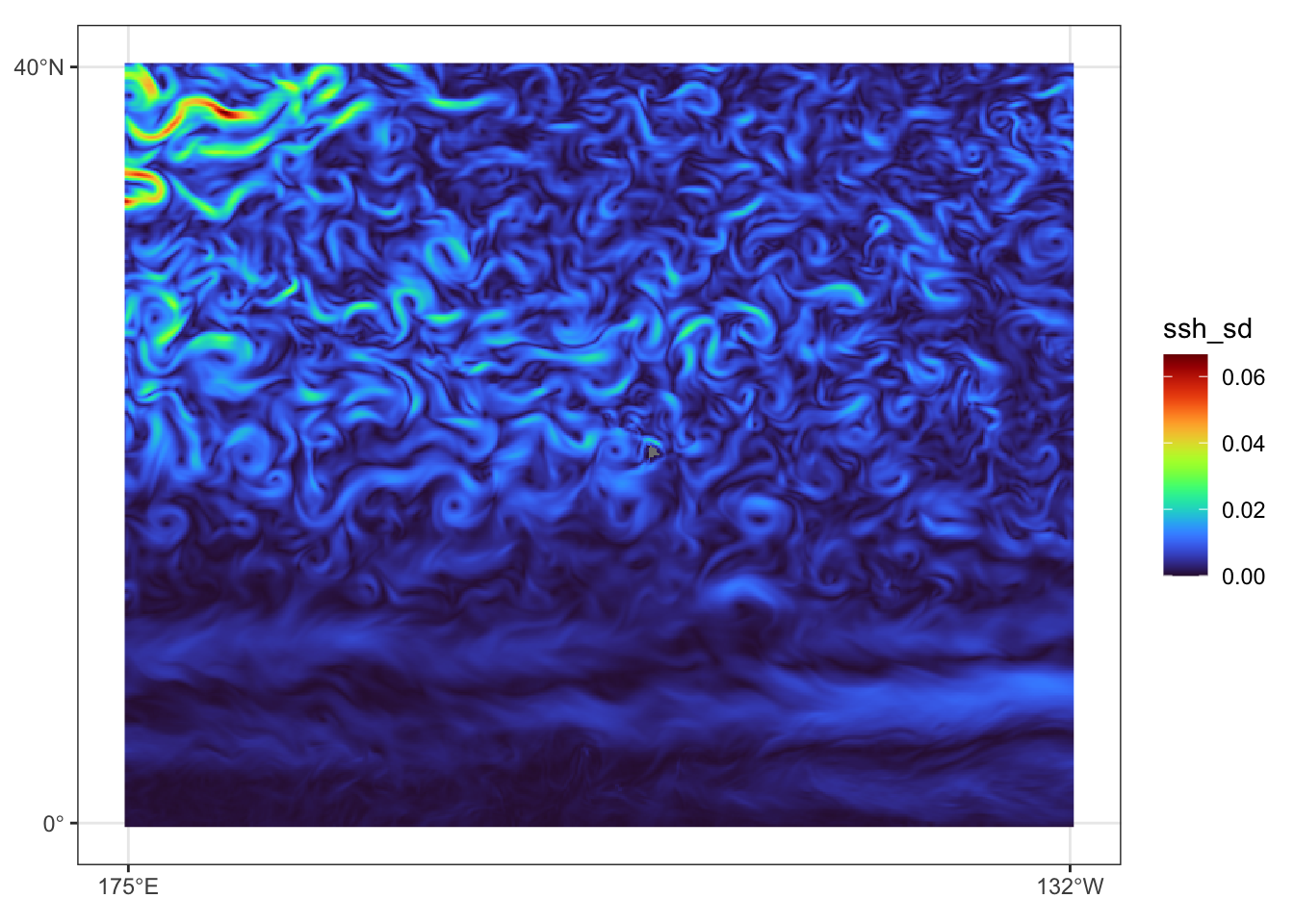
})Prediction data view for July 31, 2023