library(mgcv)
library(here)
library(tidyverse)
library(sf)
library(terra)
library(tidyterra)
library(lubridate)
library(picMaps)
library(ggspatial)
library(viridisLite)
library(viridis)
library(ggpubr)
library(mapview)
library(doFuture)
library(foreach)
local_wd <- here("_R_code","task_7_predicting_abundance")
dir.create(file.path(local_wd, "output"))
dir.create(file.path(local_wd, "output","predictions"))
env_dir <- here("_R_code","task_5_download_prediction_data","output")Predicting Spatial Abundance
A. Abundance Prediction
Packages and set up
Load model output
load(here("_R_code","task_4_fit_dsm_models","output","dsm_fit_final.RData"))Raster and storage tools
years <- read_csv(here("_R_code","task_5_download_prediction_data","copernicus_data","pred_download.csv")) %>% pull(year)
env_tmp <- rast(file.path(env_dir,"cenpac_2017_env.tif"))[[1]]
area <- rast(ext(env_tmp), resolution=res(env_tmp)); crs(area) <- crs(env_tmp)
area <- cellSize(area, unit="km")
hi_r <- picMaps::hawaii_coast() %>% rasterize(area, touches=TRUE, cover=TRUE, background=0)
hi_r <- 1-hi_r
hi_r <- ifel(hi_r==min(values(hi_r)), 0, hi_r)
cenpac_r <- picMaps::cenpac() %>% rasterize(area, touches=TRUE, cover=TRUE) * hi_r
eez_r <- picMaps::hawaii_eez() %>% rasterize(area, touches=TRUE, cover=TRUE) * hi_r
fkw_mgmt_r <- picMaps::pfkw_mgmt() %>% rasterize(area, touches=TRUE, cover=TRUE) * hi_rDetermine clamping and truncating values
fit_var <- fkw_dsm_data %>% select(sst:ssh_sd, Latitude)
clamp_df <- data.frame(var=colnames(fit_var), low=NA, high=NA, trunc_low=NA, trunc_high=NA)
for(i in 1:nrow(clamp_df)){
v <- fkw_dsm_data[[clamp_df$var[i]]]
clamp_df$low[i] <- quantile(v, 0.001)
clamp_df$high[i] <- quantile(v, 0.999)
clamp_df$trunc_low[i] <- 0.5*min(v)
clamp_df$trunc_high[i] <- 1.5*max(v)
}
save(clamp_df, file=file.path(local_wd,"output","clamp_df.RData"))Loop over years and dates to make predictions
# plan("multisession",workers=8)
N_data <- NULL
for(i in seq_along(years)){
env <- rast(file.path(env_dir, paste0("cenpac_",years[i],"_env.tif")))
times <- time(env) %>% unique() %>% sort()
for(j in seq_along(times)){
sub_idx <- (terra::time(env)==times[j])
env_j <- terra::subset(env, sub_idx)
lat <- rast(env_j, nlyr=1); values(lat) <- terra::crds(lat)[,"y"]
# clamp
for(k in 1:nlyr(env_j)){
clamp <- clamp_df[with(clamp_df, var==names(env_j)[k]),]
env_j[[k]] <- ifel(env_j[[k]]<clamp$low, clamp$low, env_j[[k]])
env_j[[k]] <- ifel(env_j[[k]]>clamp$high, clamp$high, env_j[[k]])
}
env_j$Latitude <- lat
# Predict er and gs
pred_er <- terra::predict(env_j, er_fit, type="response")
pred_gs <- exp(terra::predict(env_j, gs_fit) + 0.5*gs_fit$sig2) * gs_bias_corr
pred <- pred_er * pred_gs * area
pred <- ifel(is.na(pred), 0, pred)
N_cenpac <- as.numeric(global(pred, "sum", weights=cenpac_r, na.rm=TRUE))
N_eez <- as.numeric(global(pred, "sum", weights=eez_r, na.rm=TRUE))
N_fkw <- as.numeric(global(pred, "sum", weights=fkw_mgmt_r, na.rm=TRUE))
outfile <- paste0("N_",times[j],"_file.tif")
N_data <- bind_rows(
N_data,
data.frame(year=years[i], date=times[j], N_cenpac=N_cenpac, N_eez=N_eez,
N_fkw_mgmt=N_fkw, file=outfile
)
)
writeRaster(pred, file.path(local_wd,"output","predictions",outfile), overwrite=TRUE)
cat(j," ")
}
cat("\n\n")
}
# plan("sequential")
save(N_data, clamp_df, file=file.path(local_wd, "output","N_est_df.RData"))
N_summ <- N_data %>% group_by(year) %>%
summarize(
N_cenpac_est = round(mean(N_cenpac)),
N_cenpac_sd = round(sd(N_cenpac)),
N_eez_est = round(mean(N_eez)),
N_eez_sd = round(sd(N_eez)),
N_fkw_mgmt_est = round(mean(N_fkw_mgmt)),
N_fkw_mgmt_sd = round(sd(N_fkw_mgmt))
)
N_avg <- N_data %>% filter(year!=2017) %>%
summarize(
N_cenpac_est = round(mean(N_cenpac)),
N_cenpac_sd = round(sd(N_cenpac)),
N_eez_est = round(mean(N_eez)),
N_eez_sd = round(sd(N_eez)),
N_fkw_mgmt_est = round(mean(N_fkw_mgmt)),
N_fkw_mgmt_sd = round(sd(N_fkw_mgmt))
)
N_summ$year <- as.character(N_summ$year)
N_avg <- bind_cols(year="avg 2020-2024", N_avg)
N_summ <- bind_rows(N_avg, N_summ)
write_csv(N_summ, file=file.path(local_wd,"output","N_summ.csv"))Create yearly average rasters
pred_avg <- vector("list",length(years))
for(i in seq_along(years)){
tifs <- N_data$file[N_data$year==years[i]]
pred <- rast(file.path(local_wd, "output","predictions",tifs[1]))
for(j in 2:length(tifs)){
pred <- c(pred, rast(file.path(local_wd, "output","predictions",tifs[j])))
}
pred_avg[[i]] <- mean(pred)
}
pred_avg <- do.call(c, pred_avg)
names(pred_avg) <- years
writeRaster(pred_avg, file.path(local_wd,"output","yearly_avg_abund.tif"), overwrite=TRUE)Create abundance plots
hi <- hawaii_coast()
eez <- hawaii_eez()
fkwm <- pfkw_mgmt()
p_mean <- mean(pred_avg[[2:6]])
ppp <- vector("list",nlyr(pred_avg)+1)
ppp[[1]] <- ggplot() + geom_spatraster(data=p_mean) +
layer_spatial(hi, fill="white", color=NA) +
layer_spatial(eez, color="white", fill=NA, linetype="dashed", lwd=0.5) +
layer_spatial(fkwm, color="white", fill=NA, lwd=0.5) +
scale_fill_viridis(option="H", name="Abundance") +
scale_x_continuous(breaks=c(175, 228)) + scale_y_continuous(breaks=c(0, 40)) +
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank()) +
coord_sf(expand = FALSE) +
ggtitle("Avg. 2020-2024")
lll <- get_legend(ppp[[1]])
ppp[[1]] <- ppp[[1]] + theme(legend.position="none")
years <- names(pred_avg)
for(i in seq_along(years)){
ppp[[i+1]] <- ggplot() + geom_spatraster(data=pred_avg[[i]]) +
layer_spatial(hi, fill="white", color=NA) +
layer_spatial(eez, color="white", fill=NA, linetype="dashed", lwd=0.5) +
layer_spatial(fkwm, color="white", fill=NA, lwd=0.5) +
scale_fill_viridis(option="H", name="Abundance") +
scale_x_continuous(breaks=c(175, 228)) + scale_y_continuous(breaks=c(0, 40)) +
theme_bw() + theme(legend.position="none") +
theme(axis.ticks = element_blank()) +
theme(axis.text = element_blank(), axis.ticks = element_blank()) +
coord_sf(expand = FALSE) +
ggtitle(years[i])
}
pout <- ggarrange(ppp[[1]], ppp[[2]], ppp[[3]], ppp[[4]], ppp[[5]], ppp[[6]], ppp[[7]],lll)
ggsave(pout, file=file.path(local_wd, "output","cenpac_abund.png"), width=6.5, height=6.2)
## Assessment area
ppp <- vector("list",nlyr(pred_avg)+1)
eez_asses <- st_union(eez, fkwm) %>% st_shift_longitude()
bound <- eez_asses %>% st_buffer(100000) %>% st_shift_longitude()
pred_clip <- crop(pred_avg, vect(bound)) %>% mask(eez_asses)
pmean_clip <- crop(p_mean, vect(bound)) %>% mask(eez_asses)
ppp[[1]] <- ggplot() + geom_spatraster(data=pmean_clip) +
layer_spatial(hi, fill="black", color=NA) +
layer_spatial(eez, color="black", fill=NA, linetype="dashed", lwd=0.5) +
layer_spatial(fkwm, color="black", fill=NA, lwd=0.5) +
scale_fill_viridis(option="H", name="Abundance", na.value=NA) +
theme_bw() +
theme(axis.text = element_blank(), axis.ticks = element_blank(), panel.grid.major = element_blank()) +
coord_sf(expand = FALSE) +
ggtitle("Avg. 2020-2024")
lll <- get_legend(ppp[[1]])
ppp[[1]] <- ppp[[1]] + theme(legend.position="none")
years <- names(pred_avg)
for(i in seq_along(years)){
ppp[[i+1]] <- ggplot() + geom_spatraster(data=pred_clip[[i]]) +
layer_spatial(hi, fill="black", color=NA) +
layer_spatial(eez, color="black", fill=NA, linetype="dashed", lwd=0.5) +
layer_spatial(fkwm, color="black", fill=NA, lwd=0.5) +
scale_fill_viridis(option="H", name="Abundance", na.value=NA) +
theme_bw() + theme(legend.position="none") +
theme(axis.ticks = element_blank()) +
theme(axis.text = element_blank(), axis.ticks = element_blank(), panel.grid.major = element_blank()) +
coord_sf(expand = FALSE) +
ggtitle(years[i])
}
pout <- ggarrange(ppp[[1]], ppp[[2]], ppp[[3]], ppp[[4]], ppp[[5]], ppp[[6]], ppp[[7]],lll)
ggsave(pout, file=file.path(local_wd, "output","assess_abund.png"), width=6.5, height=4.5)Abundance prediction figures
Central Pacific study area
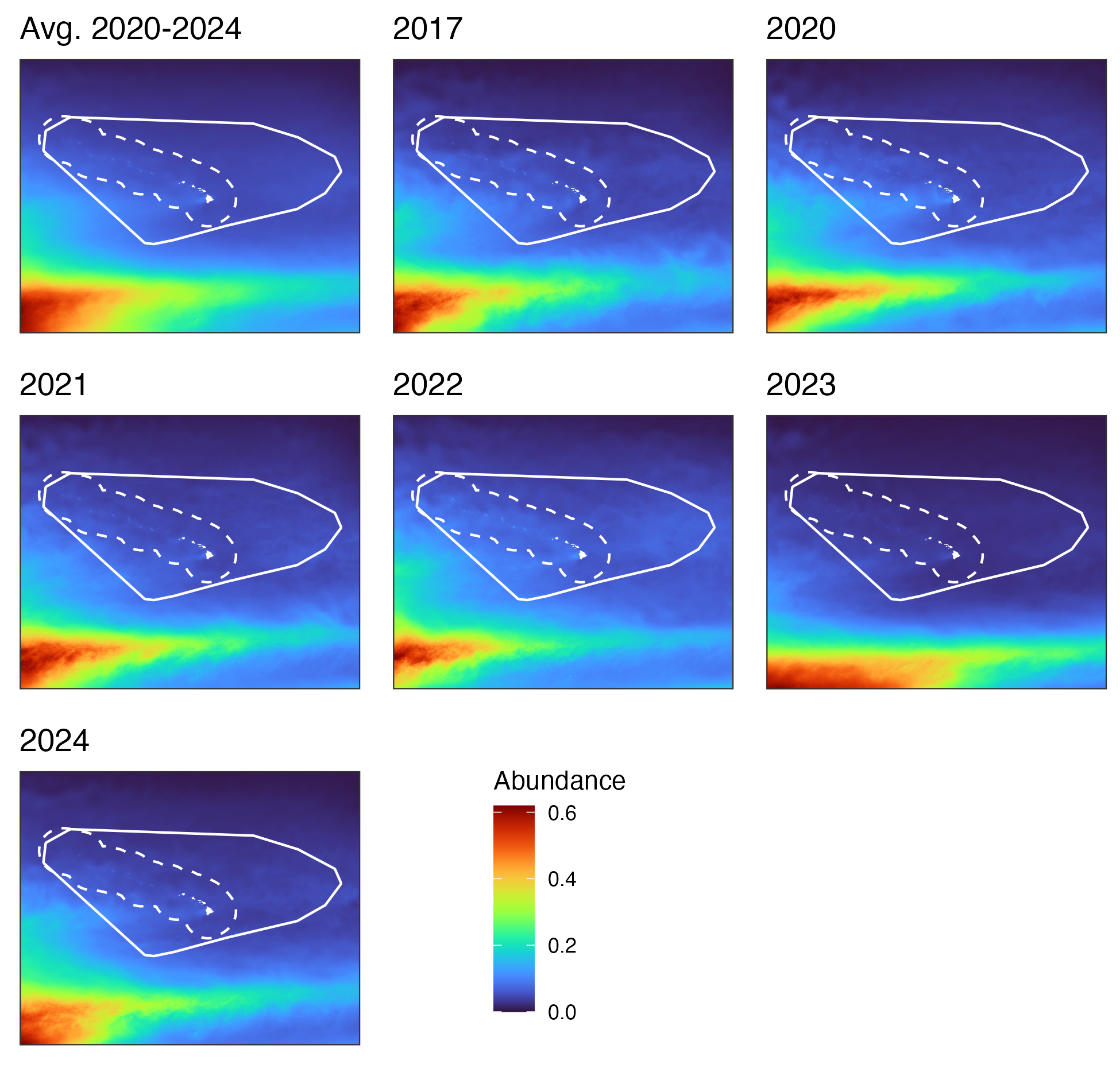
Hawaiʻi pelagic false killer whale assessment area and Hawaiian EEZ
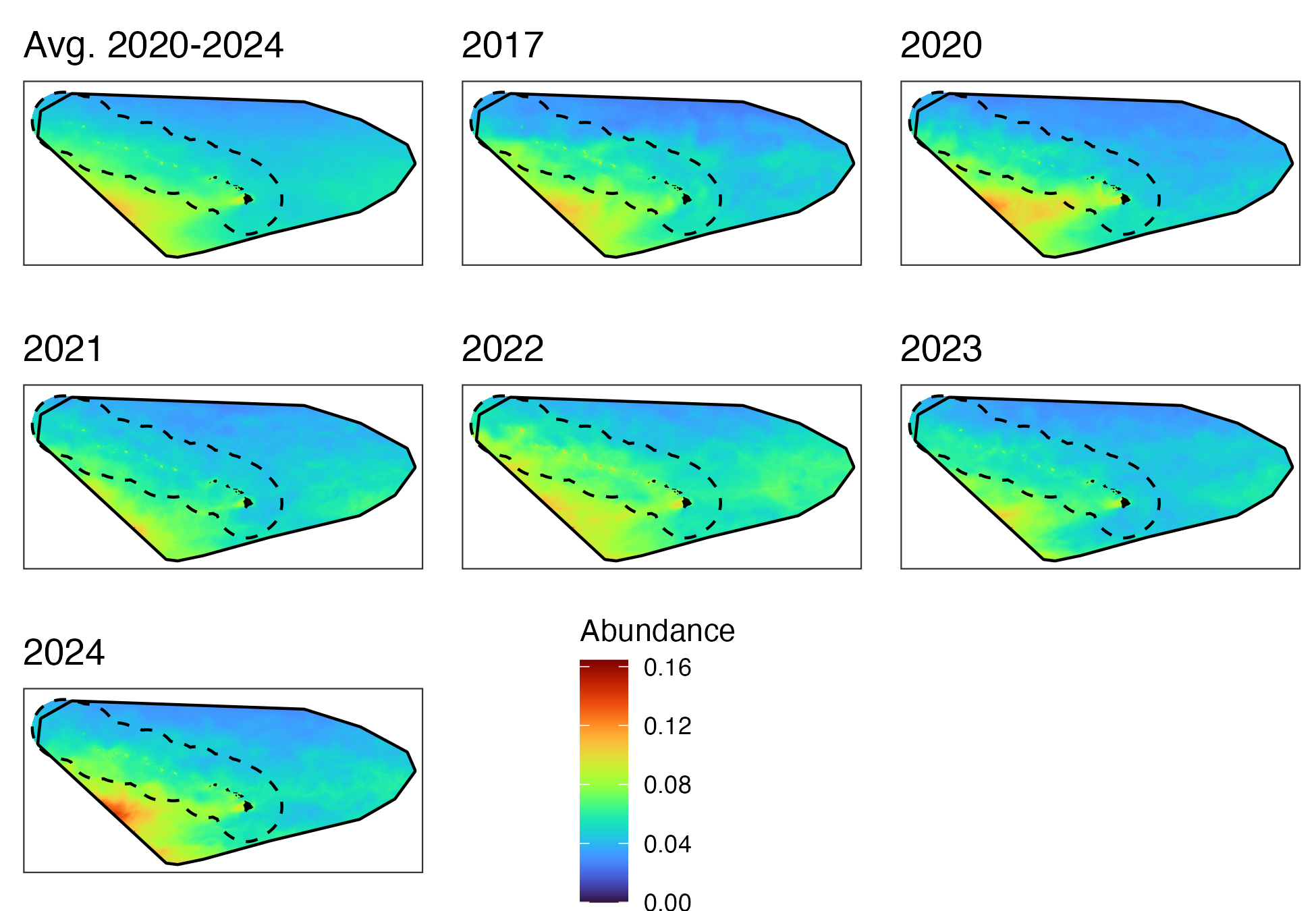
B. Model Assessment Using Previous Data (2017 and Prior)
Refit GAMs without new data (2020-2023 and 2010, cruise 1004)
fkw_dsm_data <- fkw_dsm_data %>% filter(year<=2017, Cruise!=1004)
fkw_gs <- fkw_gs %>% filter(year<=2017, Cruise!=1004)
spline2use <- "ts"
er_fit <- gam(as.list(er_fit$call)$formula,
offset = log(effort),
family = tw(),
method="REML",
data = fkw_dsm_data, weights = ProbPel)
gs_fit <- gam(as.list(gs_fit$call)$formula,
method="REML",
data = fkw_gs, weights = ProbPel)Raster and storage tools
env_tmp <- rast(file.path(env_dir,"cenpac_2017_env.tif"))[[1]]
area <- rast(ext(env_tmp), resolution=res(env_tmp)); crs(area) <- crs(env_tmp)
area <- cellSize(area, unit="km")
hi_r <- picMaps::hawaii_coast() %>% rasterize(area, touches=TRUE, cover=TRUE, background=0)
hi_r <- 1-hi_r
hi_r <- ifel(hi_r==min(values(hi_r)), 0, hi_r)
cenpac_r <- picMaps::cenpac() %>% rasterize(area, touches=TRUE, cover=TRUE)
cenpac_r <- cenpac_r * hi_r
eez_r <- picMaps::hawaii_eez() %>% rasterize(area, touches=TRUE, cover=TRUE)
fkw_mgmt_r <- picMaps::pfkw_mgmt() %>% rasterize(area, touches=TRUE, cover=TRUE)Determine clamping and truncating values
fit_var <- fkw_dsm_data %>% select(sst:ssh_sd, Latitude)
clamp_df <- data.frame(var=colnames(fit_var), low=NA, high=NA, trunc_low=NA, trunc_high=NA)
for(i in 1:nrow(clamp_df)){
v <- fkw_dsm_data[[clamp_df$var[i]]]
clamp_df$low[i] <- quantile(v, 0.001)
clamp_df$high[i] <- quantile(v, 0.999)
clamp_df$trunc_low[i] <- 0.5*min(v)
clamp_df$trunc_high[i] <- 1.5*max(v)
}Loop over years and parameters to make predictions
# plan("multisession",workers=8)
N_data <- NULL
env <- rast(file.path(env_dir, "cenpac_2017_env.tif"))
times <- time(env) %>% unique() %>% sort()
for(j in seq_along(times)){
sub_idx <- (terra::time(env)==times[j])
env_j <- terra::subset(env, sub_idx)
lat <- rast(env_j, nlyr=1); values(lat) <- terra::crds(lat)[,"y"]
# clamp
for(k in 1:nlyr(env_j)){
clamp <- clamp_df[with(clamp_df, var==names(env_j)[k]),]
env_j[[k]] <- ifel(env_j[[k]]<clamp$low, clamp$low, env_j[[k]])
env_j[[k]] <- ifel(env_j[[k]]>clamp$high, clamp$high, env_j[[k]])
}
env_j$Latitude <- lat
# Predict er and gs
pred_er <- terra::predict(env_j, er_fit, type="response")
pred_gs <- exp(terra::predict(env_j, gs_fit) + 0.5*gs_fit$sig2) * gs_bias_corr
pred <- pred_er * pred_gs * area * cenpac_r
pred <- ifel(is.na(pred), 0, pred)
N_cenpac <- as.numeric(global(pred, "sum", na.rm=TRUE))
N_eez <- as.numeric(global(pred, "sum", weights=eez_r, na.rm=TRUE))
N_fkw <- as.numeric(global(pred, "sum", weights=fkw_mgmt_r, na.rm=TRUE))
outfile <- paste0("N_",times[j],"_file.tif")
N_data <- bind_rows(
N_data,
data.frame(year=2017, date=times[j], N_cenpac=N_cenpac, N_eez=N_eez,
N_fkw_mgmt=N_fkw, file=outfile
)
)
cat(j," ")
}
save(N_data, clamp_df, file=file.path(local_wd, "output","previous_data","N_est_df.RData"))
N_summ <- N_data %>% group_by(year) %>%
summarize(
N_cenpac_est = round(mean(N_cenpac)),
N_cenpac_sd = round(sd(N_cenpac)),
N_eez_est = round(mean(N_eez)),
N_eez_sd = round(sd(N_eez)),
N_fkw_mgmt_est = round(mean(N_fkw_mgmt)),
N_fkw_mgmt_sd = round(sd(N_fkw_mgmt))
)
write_csv(N_summ, file=file.path(local_wd,"output","previous_data","N_summ.csv"))Abundance prediction results using only previous data
# A tibble: 1 × 7
year N_cenpac_est N_cenpac_sd N_eez_est N_eez_sd N_fkw_mgmt_est N_fkw_mgmt_sd
<dbl> <dbl> <dbl> <dbl> <dbl> <dbl> <dbl>
1 2017 29171 5834 2268 819 5335 1779